- #Download cytoscape how to#
- #Download cytoscape install#
- #Download cytoscape update#
- #Download cytoscape code#
- #Download cytoscape license#
Gallery Dynamically expand elementsĬode | Demo Interactively update stylesheetĬode | Demo Automatically generate interactive phylogeny treesįor an extended gallery, visit the demos' readme. Overall, the Gephi interface is a bit more intuitive.
#Download cytoscape how to#
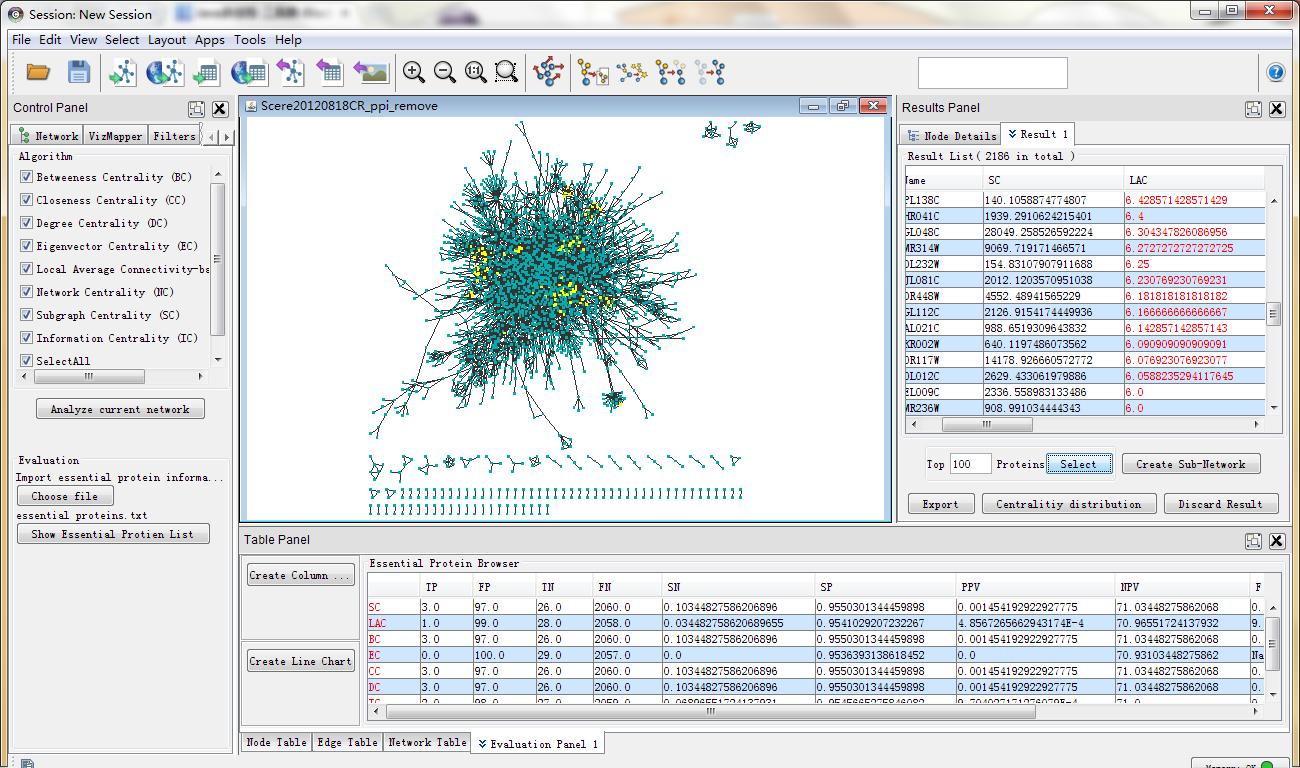
4) how to calculate and look up metrics on both platforms. 3) how to adjust settings for layout algorithms. Download Cytoscape - View and analyze molecular interaction networks using a comprehensive application that puts at your disposal a wide number of annotation methods. 2) how to create and style network visualizations. The Pull Request and Issue Templates were inspired from the In this introductory tutorial to network visualization in Cytoscape and Gephi, we essentially covered : 1) how to upload a dataset.
This library would not have been possible without their massive work! Huge thanks to the Cytoscape Consortium and the Cytoscape.js team for their contribution in making such a complete API for creating interactive networks. Licenseĭash, Cytoscape.js and Dash Cytoscape are licensed under MIT. Instructions on how to run tests are given in CONTRIBUTING.md.
#Download cytoscape code#
Make sure that you have read and understood our code of conduct, then head over to CONTRIBUTING to get started. To learn more about the core Dash components and how to use callbacks, view the Dash documentation.įor supplementary information about the underlying Javascript API, view the Cytoscape.js documentation. You can also use the component reference for a complete and concise specification of the API. It contains useful examples, functioning code, and is fully interactive.
#Download cytoscape install#
You can manually download and install latest version of. The Dash Cytoscape User Guide contains everything you need to know about the library. Cytoscape installer automatically downloads a suitable Java 8 if none is available on your workstation. Install the library using devtools: devtools::install_github("plotly/dash-cytoscape")Ĭreate the following example inside an app.R file: library ( dash ) library ( dashHtmlComponents ) library ( dashCytoscape ) app <- Dash $ new () app $ layout ( htmlDiv ( list ( cytoCytoscape ( id = 'cytoscape-two-nodes', layout = list ( 'name' = 'preset' ), style = list ( 'width' = '100%', 'height' = '400px' ), elements = list ( list ( 'data' = list ( 'id' = 'one', 'label' = 'Node 1' ), 'position' = list ( 'x' = 75, 'y' = 75 )), list ( 'data' = list ( 'id' = 'two', 'label' = 'Node 2' ), 'position' = list ( 'x' = 200, 'y' = 200 )), list ( 'data' = list ( 'source' = 'one', 'target' = 'two' )) ) ) ) ) ) app $ run_server () Documentation Getting Started in R Prerequisites install.packages ( c ( "devtools", "dash" )) Usage Div ()Ĭalling cyto.load_extra_layouts() also enables generating SVG images. Use the cyto.load_extra_layouts() function to get started: import dash import dash_cytoscape as cyto import dash_html_components as html cyto. Install the library using pip: pip install dash-cytoscapeĬreate the following example inside an app.py file: import dash import dash_cytoscape as cyto import dash_html_components as html app = dash. If you want to install the latest versions, check out the Dash docs on installation. Designed for users first, for both frontfacing. Used in commercial projects and open-source projects in production.
#Download cytoscape license#
Permissive open source license (MIT) for the core Cytoscape.js library and all first-party extensions. A fully featured graph library written in pure JS. Make sure that dash and its dependent libraries are correctly installed: pip install dash Graph theory library for visualization and analysis. To run the tutorial you will need to install RC圓: if(!"RC圓" %in% installed.A Dash component library for creating interactive and customizable networks in Python, wrapped around Cytoscape.js. More information on the database contents, statistics and attributes is available here. Additionally, several original metrics are provided to assist the prioritization of genotype–phenotype relationships. The data in the platform are homogeneously annotated with controlled vocabularies and community-driven ontologies. DisGeNET integrates data from expert curated repositories, GWAS catalogues, animal models and the scientific literature. A tutorial on how to use the DisGeNET Cytoscape App from Cytoscape is available here.ĭisGeNET is a discovery platform containing one of the largest publicly available collections of genes and variants associated with human diseases. This tutorial will teach you how to use the DisGeNET Cytoscape App to retrieve data from DisGeNET using the R programming language.
